|
In the case of sfGFP purification, cell lysate was heat-treated at 80☌ for 15 min before centrifugation. Harvested cells were sonicated in buffer (20 mM Tris-HCl, pH 7.4, 1 mM DTT, a Complete protease inhibitor mixture tablet (Roche Applied Science)) and centrifuged. Proteins were expressed in Escherichia coli BL21(DE3) at 20☌ for 20 hr. A206K mutations (monomer propensity) and S208F/V224L mutations (dimer propensity) were also introduced. A cysteine residue was introduced at the N-terminus of CFP (Cys-CFP) and at the 173 position of YFP (Cys-YFP), taking into consideration their orientations in the dimer. We used Cerulean and Venus as improved CFP and YFP. This increases local concentrations of the proteins and will enable association of proteins with very weak interactions.Įxpression plasmid pET21c-His 6-sfGFP-Cys was generated from pET21c-wild-type GFP by site-directed mutagenesis using the PrimeSTAR mutagenesis basal kit (Takara, Japan). Here we report a procedure to align several different proteins, all connected to a single flexible DNA backbone. For effective formation of complex, proteins should be linked with high flexibility enough to allow proteins to move and turn freely to associate with each other. Fused protein has structural restriction and is often difficult to express in a soluble form. Chemical cross-linking has a limited reactivity and is not easy to control the number of cross-linked proteins. However, these methods have some limitations. To study such complex, component proteins of the complex are chemically cross-linked or genetically fused as a single polypeptide to prevent the complex from dissociation. In many cases, however, interactions are transient or weak, producing a complex that is difficult to isolate. When the interactions are strong, proteins associate into a stable homo- or hetero-oligomer complex that can be isolated relatively easily. Protein-protein interactions play a critical role in numerous biological processes, and understanding the nature of each interaction is of central importance in biology and biotechnology. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.Ĭompeting interests: The authors have declared that no competing interests exist. Y.) from the Ministry of Education, Culture, Sports, Science and Technology, Japan. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.įunding: This work was supported by a Grant-in-Aid for Scientific Research on Priority Areas (No. Received: JAccepted: NovemPublished: December 26, 2012Ĭopyright: © 2012 Nojima et al. School of Chemistry and Biochemistry, Georgia Institute of Technology, Atlanta, Georgia 30332-0400, USA.Citation: Nojima T, Konno H, Kodera N, Seio K, Taguchi H, Yoshida M (2012) Nano-Scale Alignment of Proteins on a Flexible DNA Backbone. This bending may be related to the direct coordination of a sodium cation by a DNA base, with unprecedented inner-shell (direct) coordination of penta-hydrated sodium at the O6 atom of a guanine. Electrostatic forces appear to induce modest DNA bending into the major groove. TriplatinNC extends along the phosphate backbone, in a mode of binding we call "Backbone Tracking" and spans the minor groove in a mode of binding we call "Groove Spanning". The high repetition and geometric regularity of the motif suggests that this type of Pt(II) center can be developed as a modular nucleic acid binding device with general utility. The interaction appears to prefer O2P over O1P atoms (frequency of interaction is O2P > O1P, base and sugar oxygens > N). The geometry is conserved among the 8 observed phosphate clamps in this structure. The three square-planar tetra-am(m)ine Pt(II) coordination units form bidentate N.O.N complexes with OP atoms, in a motif we call the Phosphate Clamp. 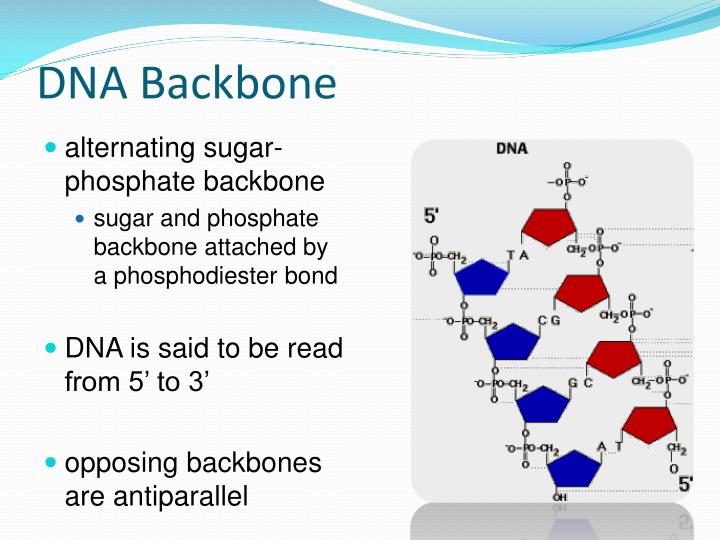
Instead, it binds to phosphate oxygen atoms and thus associates with the backbone. TriplatinNC does not intercalate nor does it bind in either groove. TriplatinNC is a multifunctional DNA ligand, with three cationic Pt(II) centers, and directional hydrogen bonding functionalities, linked by flexible hydrophobic segments, but without the potential for covalent interaction. 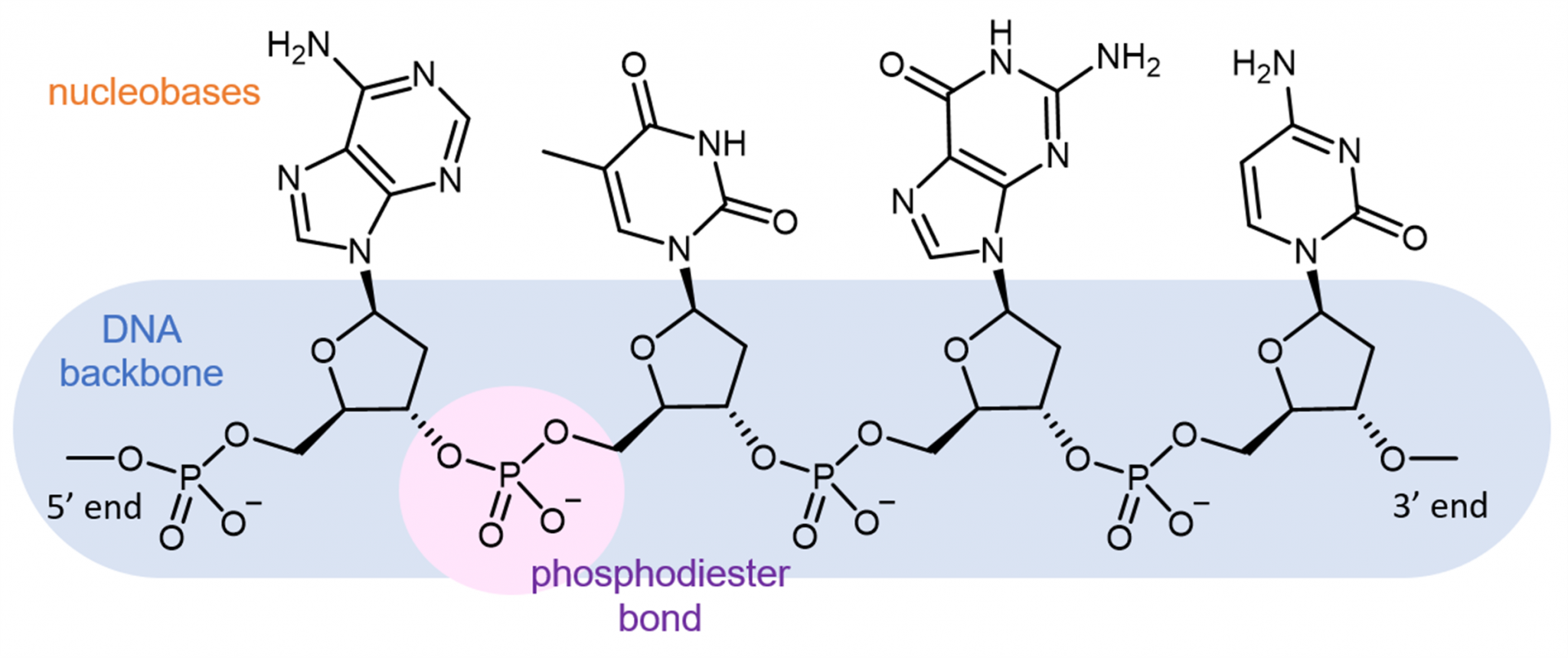
We describe a 1.2 A X-ray structure of a double-stranded B-DNA dodecamer (the Dickerson Dodecamer, DDD, 2) associated with a cytotoxic platinum(II) complex, (TriplatinNC).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
